On target (Last updated: 2025-06-16)

A beginners introduction to getting started with the targets package in R
Here are some notes on getting started with {targets}.
The project I am working on involves several different reports, each at least 30 pages, and each with about 20 plots and 20 tables per document.
As well as a myriad of functions, I had 7 very large R scripts doing the data munging and processing.
I thought they were well ordered, but I had to burn everything down a couple of times and it was quite nerve wracking building it back up, as there were a few inter-related parts scattered about.
The thought of adding additional phases of the project to this code base made me uncomfortable. I decided I needed to learn {targets} to ensure this project can be reproducible a few years down the line.
The package comes with extensive documentation, but here are some edited highlights and explainers
If you don’t know what targets does - it keeps track of the objects you create, and the relationships between those objects. So if you have a file that feeds into a function, and the file updates, then the function needs to be run again. You don’t need to keep track of that in your head, {targets} does the work for you and produces a wonderful network plot showing the current status.
For example - here is a very zoomed out view of all my targets. It’s hard to tell, but quite a lot are now out of date - as seen by the blue colour
Here I’ve zoomed in, with particular focus on the localities target, which acts as an input to many other downstream targets.
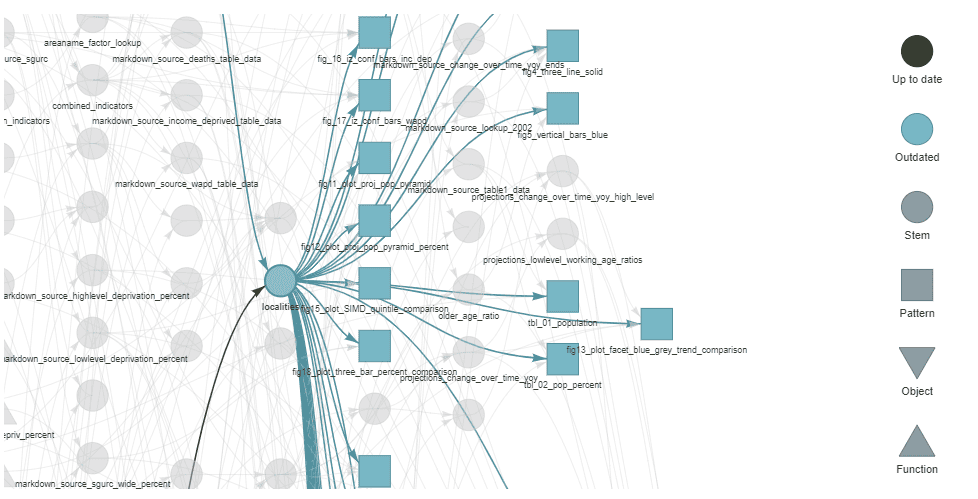
The next time tar_make is run, the code that updates these functions will run, and everything else will be skipped. There is no way, having broken everything down into small functions, that I could track all this manually.
Note - I’m using dataframe here as a generic term for data.frame, tibble, data.table, or whatever else you might be using.
The name of your target matters - if you have a function that returns a dataframe, and you want that dataframe to have a specific name, then that’s the name you should give your target. E.G. if you want your function to return a dataframe called ‘sales_data’, then, that’s the name of the target.
Being specific with your target names is very important if the result of one target, e,g. a dataframe, is used as an input into another function.
By default , a target will be saved as an .RDS file. This caused me some grief initially, because my functions (written before I knew targets) were saving the outputs as RDS files so there was a bit of double handling going on. So not only did I have badly named targets, I was saving them in an output folder, and unbeknownst to me (because I didn’t RTFM), they were also being saved elsewhere. I was then passing these .RDS filenames as names for other targets, and generally in a total mess. Don’t do that.
Targets are automatically saved in “/targets/objects/your_target_name”. They don’t show as having an .RDS extension, but unless you specify otherwise, that’s what they will be (other file formats are availble, but .RDS works fine for me)
If you want to track a CSV file, or spreadsheet, or perhaps you are saving a ggplot2 plot as a PNG file, then you need to specify that the target is a “file”
For example, here I’m tracking a spreadsheet which has a list of desired indicators. If the file changes in any way, then anything that depends on this will become outdated, and {targets} will know to update those parts of the pipeline
tar_target(profile2_adult_indicators,# target name "./01-inputs/profile2_adultindicators.xlsx", # command format = "file") # target file format However, to be very clear, if you are producing a dataframe for use in other targets, then don’t specify that as a “file”
You need to use the target names as arguments in any downstream funtions that depend on them
For example, if you have a dataframe called
change_over_time, another calledlookup, andplot_three_solid_linesis a function that uses them both, then you probably want another target specified like this (wherefig_three_solid_linesis the target name)tar_target(fig_three_solid_lines, # target plot_three_solid_lines(.df1 = change_over_time, # function as command .df2 = lookup), #existing targets as inputs format = "file") # format of target outputIf you have a large script that generates several objects, you’re going to need to break that down into functions so that one target is returned per function. It seems a lot of work, but its worth it.
In general, you will use tar_manifest, tar_vis_network, and tar_make the most.
tar_manifest creates a table of the targets and their inputs and contains lots of info that will help you check that everything is working. If you run tar_make, and your pipeline doesn’t work as expected, you need to run tar_manifest and examine the output in detail (you’ll probably want to pipe the output straight into dplyr::View(.)
You might also want to use tar_invalidate with a specific target to ensure any changes you make, e.g., as a result of a function change, are picked up prior to running tar_make.
You can also use tar_destroy and set the option to “all” to completely burn everything down and rebuild it. Probably not something to use on a Friday afternoon, unless you’re very confident in your pipeline, or you simply live for the danger.
- iterating over plots. In my plotting functions, I like to have the option to print to screen and/ or save to disk. Prior to using {targets}, I used
purrr::walk2()to map over council and area values and save plots to disk. With {targets} and {tarchetypes}, it’s possible to achieve the same thing, without relying on purrr
tar_target( fig_three_bar_comparison, #target name plot_three_bar_comparison(df = combined_populations, # command council = localities$area, areaval = localities$areaname), pattern = map(localities), # localities = a 2 column df with area & areaname iteration = "list", format = "file" # format ) This code maps over the area and areanamecolumns in my localities dataframe, and creates a plot for each combination, using the plot_three_bar_comparison function, with existing target combined_populations as an input. The target name is fig_three_bar_comparison
Here are the results of this bit of code:
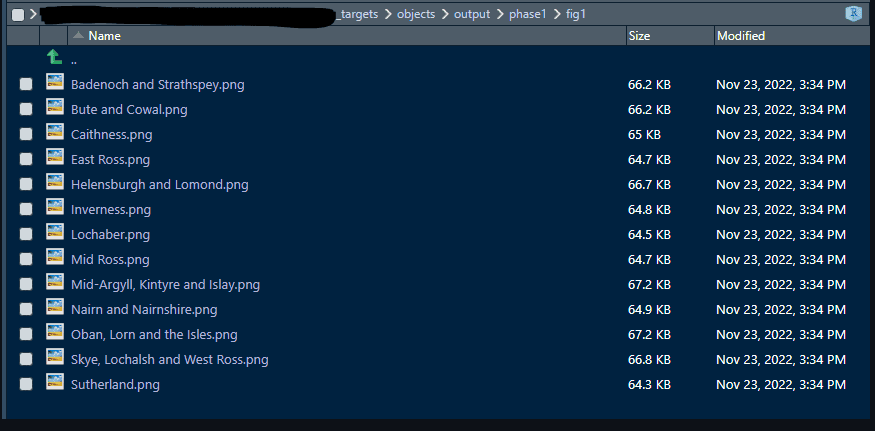
This is using dynamic branching . Static branching is also available, and I should probably have used it, as I know what I want my file names to be. I’m using static branching to generate each Word document with tar_render. This involves creating a tibble with the column values to map over, and an output vector. That may be the topic of a future post.
I have many of these functions, and as a result I already have over 500 individual targets, from original source spreadsheets and CSVs to plots and documents.
I am much happier now about the foundations of the project. I had a draft phase 2 document up and running in a couple of days - and this is a much larger document with even more tables and plots. Combined with {renv} and git, we are in a good place for our first Reproducible Analytic Pipeline.